Antibiotics are complex chemical compounds that destroy pathogenic microbes causing dangerous diseases. Penicillin was first discovered antibacterial drug widely used in medicine. Scientists have discovered hundreds of different antibiotics after penicillin, but evolution is an active process: microbes have developed a possibility to destroy antibiotics causing serious problems in treatment. How did penicillin save millions of people? Did it really become a trigger for the evolution of microbes?
While working with bacterial cultures of Staphylococcus, British microbiologist Alexander Fleming noticed that essence evolved by mold fungi Penicillium inhibits the growth of bacteria. In 1929, Fleming called this substance penicillin but he couldn’t extract it. 10 years after Fleming’s discovery, another British scientists, Ernst Chain and Howard Florey, managed to separate penicillin and introduced a possibility of its use in treating diseases. In nature, Penicillium grows on all possible biodegradable substances including foods and is toxic for human health.
Penicillin was the first antibiotic – antibacterial drug produced by microorganisms. For this research, Fleming, Chain, and Florey received the Nobel Prize in 1945. After penicillin implementation, humanity started cherishing hopes that it would tackle with all bacterial diseases. However, in a short time, there appeared antibiotic-resistant species of bacteria. Initially, less than 3% species of Staphylococcus aureus bacteria showed resistance, but today their quantity has risen up to 90%. So you ask which biological peculiarities make bacteria obtain resistance so quickly?
Antibacterial efficiency of penicillin is determined by its capability to inhibit cellular wall synthesis. Bacterial walls contain unique macromolecules – peptidoglycans, which are formed of remnants of N-acetamidodeoxyglucose (NAG) and N-acetylmuramic acids. These carbohydrate remnants are connected with protein which provides stiffness and firmness of cellular wall. Penicillin as antibiotic reduces ferment activity that catalyzes the formation of the protein binders, destroying the cell wall and leading to bacteria destruction. These ferments are called penicillin-binding protein. Penicillin and penicillin-like antibiotics are called ß-lactams.
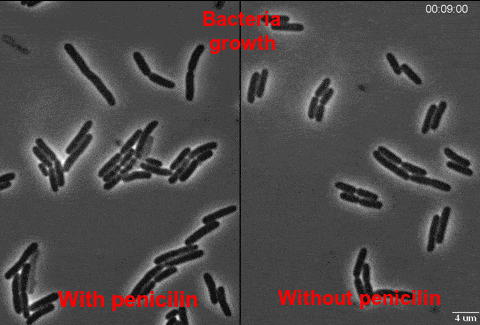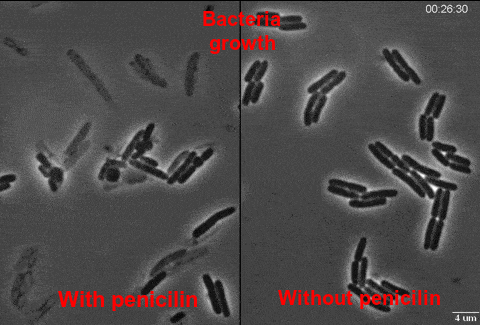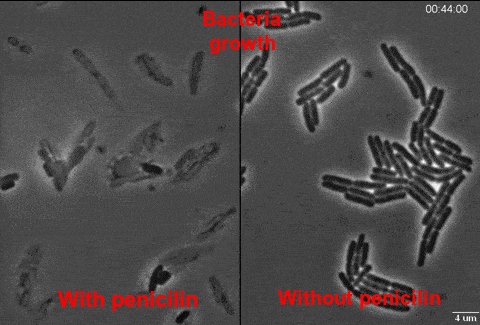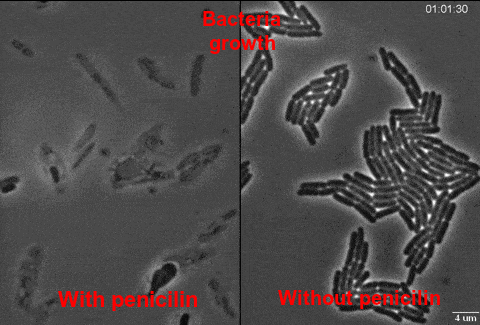
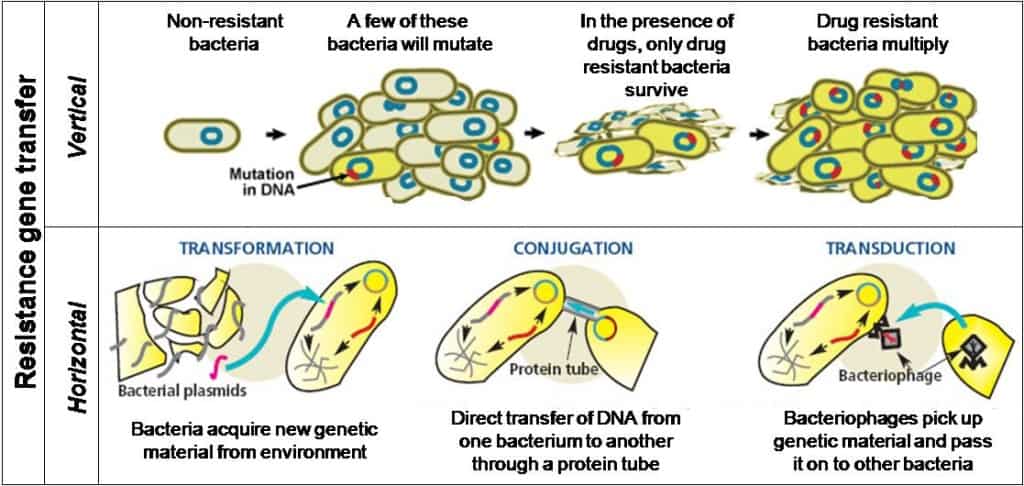
After penicillin started to be used in big amounts, lots of bacteria became insensitive and resistant. Some of the resistant bacteria synthesize penicillinase – an enzyme that destroys penicillin. In other resistant bacteria appeared mutations in genes of penicillin-binding protein, which reduced their possibility to bind protein. It means that penicillin lost its ability to inhibit the synthesis of cell wall. Under antibiotic, sensible bacteria dies out, but resistant ones stay alive to multiply and replenish. Consequently, there appear selective conditions for bacteria which have genes of resistance. This means of information transportation is called vertical. Except for bacterial “chromosome” (nucleoid), genetic bacteria apparatus is represented by plasmids – extrachromosomal ring molecules of DNA, which can be transferred from one bacterium to another (which is horizontal evolution) increasing the number of resistant bacteria. Another ability to move these genes between microbes are bacteriophages – viruses of bacteria that can transfer antibiotic resistance gene. Interestingly, microbes containing antibiotic resistance gene can be found everywhere on the Earth, even in the oceans and arctic snow.
Until recently, pneumonia agent, Streptococcus pneumonia, was susceptible to penicillin. However, there appeared resistant bacteria and mutations in genes coding penicillin-binding proteins reduced their capability to be susceptible to drugs. Moreover, these mutated genes came from other species of Streptococcus. Almost all species of Staphylococcus aureus (bacteria causing skin infections, pneumonia, food poisoning) became resistant to penicillin within last 50 years. Not long ago, these infections could be treated with antibiotics of a penicillin type (methicillin), which couldn’t be ruined due to their chemical modifications. After all, bacteria mutated again starting to synthesize penicillin-binding proteins with low penicillin contiguity. Now Methicillin-resistant Staphylococcus aureus (MRSA) is the most frequently identified antibiotic-resistant pathogen in the US hospitals.

Selective conditions for resistant microbes lead to their uncontrolled growth and appearance of different forms of diseases. Each year in the United States, at least 2 million people acquire serious infections with antibiotic-resistant strains of bacteria. That’s why it is important to determine if the disease is caused by antibiotic-resistant bacteria strain. In general, it can take few days to determine this feature, though this information is crucial for immediate treatment. Recently, a group of scientists from Arizona State University developed a technique to sort antibiotic-resistant bacteria in several minutes. Researchers have used microscale electric field gradients to identify different bacteria strains.
Antibiotic discovery happened to be a great benefit for humanity that saved millions of lives. But bacteria mutate and develop. The natural process of evolution provides new adaptations for bacteria. Quick division and capability of horizontal gene transfer in bacteria are perfect conditions for rapid spread of genes and creation of new species. The problem of resistance is of high concern among scientists – if all microbes became resistant, we would be left with no treatment and return to pre-antibiotic era. What is more, patients themselves are responsible for the resistance of bacteria to antibiotic in their own organisms. Improper use of antibiotic without doctor’s prescription, overuse or underuse are the factors due to which we create selective conditions for the life of resistant bacteria and encourage their spread.